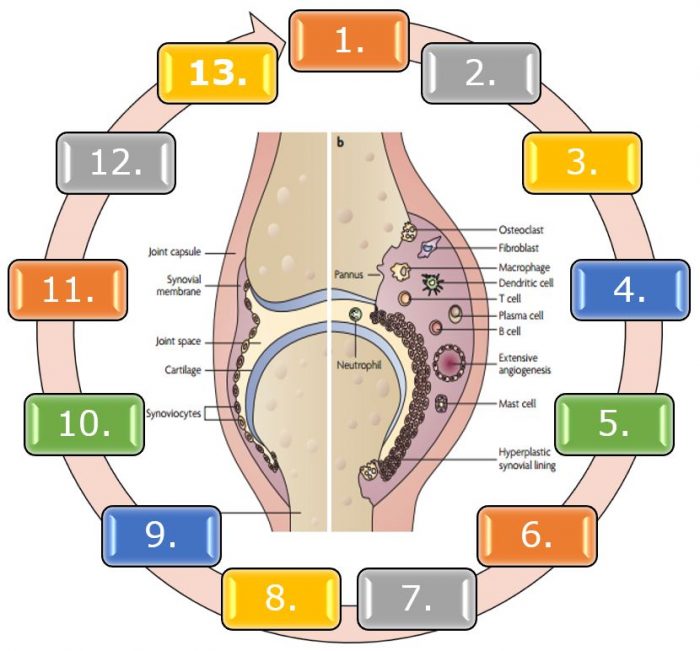
1. Exposure to the stressors (hypothalamus hypophysisadrenal axis and sympathetic nervous system), 2. Changes in microbione (gut, mouth); MAMPS/DAMPS/PAMPS activation of TLR and NOD, 4. Production of MIF, 5. Activation of NF –kB, 6.Inflammasome activation, 7. Cellular response (Th17, macrophage,EPC), 8. Hypoxia: production of HIF, 9. Angiogenesis: production of VEGF, 10. Bone and cartilage damage, 11. Failure of negative regulators (GS,AMPK,Treg,Breg), 12. Failure of apoptosis/autophagy of inflammatory cells, 13. Chronical synovitis
Project leader (PI): prof. dr. sc. Miroslav Harjaček
Host institution: University of Zagreb School of Medicine, Zagreb, Croatia
Project duration: 48 months (01.04.2017.– 01.04.2021.)
Project funding: 897.925,76 Kn
TEAM MEMBERS
1.Dr. sc. Lovro Lamot, M.D. (University of Zagreb School of Medicine)
2. Prof. dr. sc. Vedran Katavić, M.D. (University of Zagreb School of Medicine)
3. Dr. sc. Andrea Tešija Kuna (Clinical Hospital Center “Sestre Milosrdnice”)
4. Dr. sc. Danica Vidović, D.M.D. (University Hospital Centre Zagreb)
5. Dr. sc. Suzana Ožanić Bulić, M.D. (Clinical Hospital Center “Sestre Milosrdnice”)
6. Prim. Mandica Vidović, M.D. (Clinical Hospital Center “Sestre Milosrdnice)
7. Dr. sc. Mario Cindrić (Ruđer Bošković Institute)
8. Prof. dr. sc. Fran Borovečki, M.D. (University of Zagreb School of Medicine)
9. Dr. sc. Kristina Gotovac (University of Zagreb School of Medicine)
10. Domagoj Buljan, M.D. (University of Zagreb School of Medicine)
11. Ivana Radoš, M.D. (Clinical Hospital Center “Sestre Milosrdnice)
PROJECT SUMMARY AND OBJECTIVES
Juvenile idiopathic arthritis (JIA) is the most common childhood rheumatic disease. The early diagnosis of new-onset JIA has become a major objective for pediatric rheumatologists in order to identify a management strategy able to change the natural history of the disease and to prevent joint damage and functional impairment. The term undifferentiated arthritis (UA) is applied to the most common type of arthritis at the early stage when, in the absence of current recommended diagnostic criteria, it cannot be classified into the clinical subtypes of JIA. Patients with UA may progress towards JIA; however in some cases arthritis may completely resolve. Many studies has shown important role of immune system dysregulation influenced by genetic and environmental factors, but we are still far from having a clear picture of the molecular network that predisposes a child to develop the disease, to worsen the symptoms, or to successfully respond to a specific treatment. The goal of this project is to identify various epigenetic, protein and dysbiotic biomarkers in blood, stool, saliva and synovial fluid of 17 UA and 33 JIA patients, along with the value of power-doppler ultrasound (PDUS) and clinical assessment tools in the prediction of disease evolution. Therefore, we will analyze epigenetic modifications (methilatyon and histone modifications) of transcription factors and phosphatases with a known role in disease development, as well as microRNA expression profiles. Furthermore, to elucidate the possible role of other proteins, proteome will be analyzed by mass spectrometry, and selected proteins will be confirmed by complementary methods. The results of this comprehensive project will provide new insights into complex pathophysiology of JIA and enable us to use epigenetic, protein and dysbiotic biomarkers for better diagnosis, classification and treatment response prediction. In addition, results of the project may help to identify potentially new therapeutic targets.